Trusted by top biopharma companies
Thousands of med chemists from global drug discovery companies, including 50% of the top biopharma, trust StarDrop to accelerate time from hit to candidate.
Trial StarDrop



















How does StarDrop support your hit-to-candidate process?
Real-time collaboration
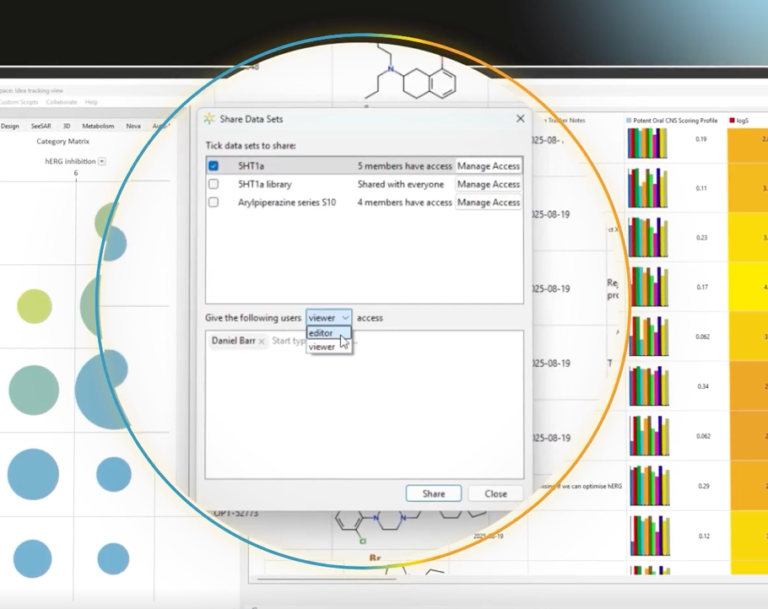
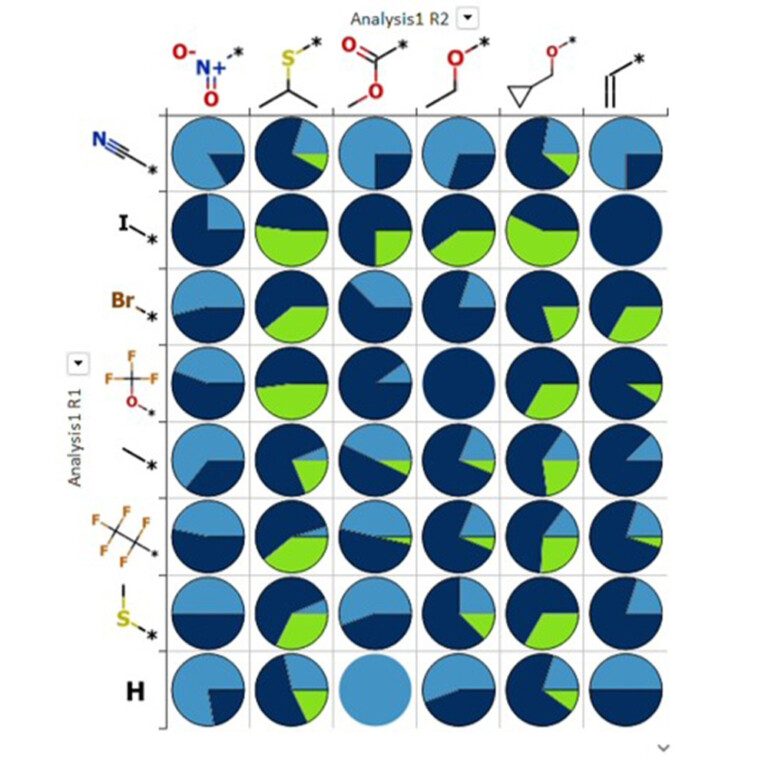
Easy SAR analysis

Clear data visualisation
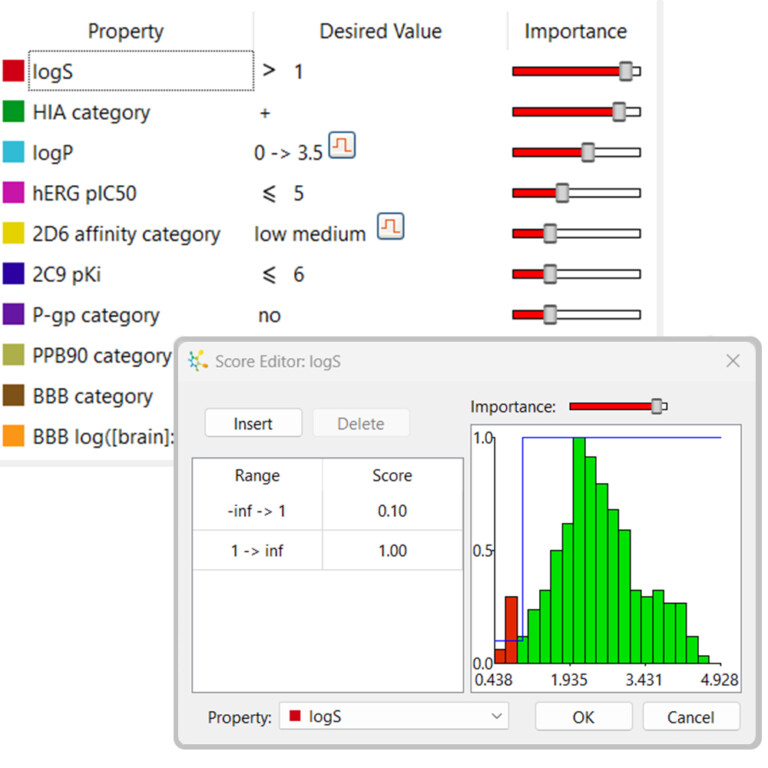
Multi-parameter optimisation
Intuitive Card View®
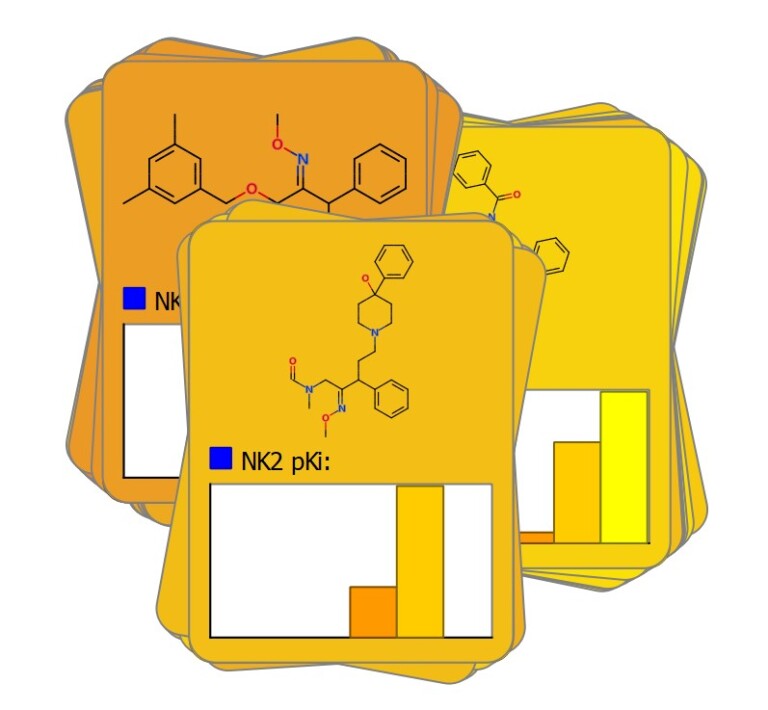
Dynamic Glowing Molecule™

Seamless AI insights
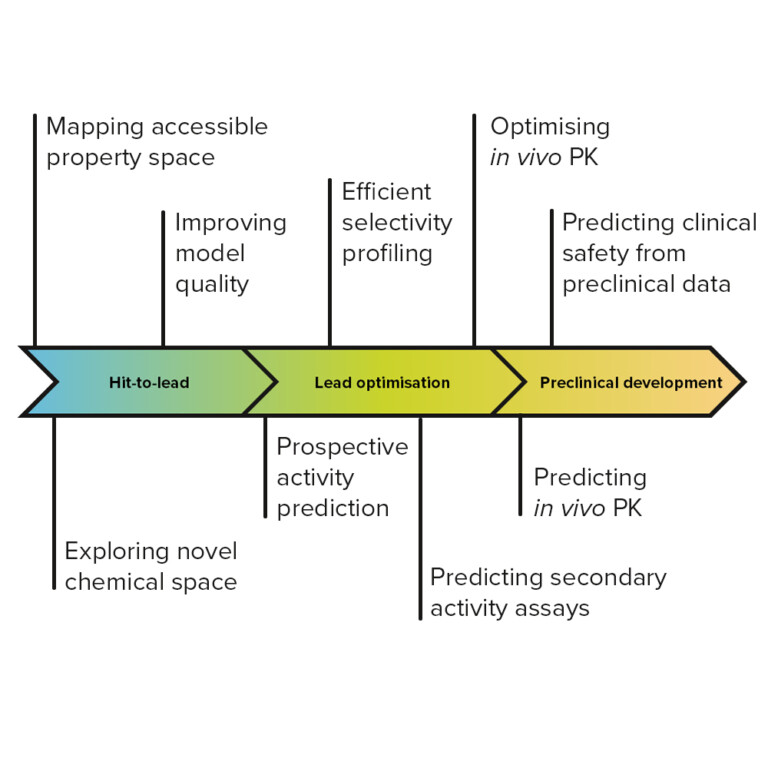
Tailored to you
Access a huge range of customisation and integration options to build the workflow you need. Plug-in modules for in silico modelling, generative chemistry or 3D design; collaborative drug design extensions; in-house and external database integration. Whatever you need for your workflow, StarDrop can help.
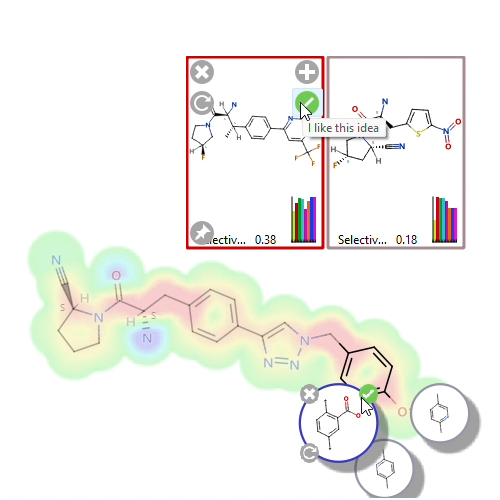
We work with thousands of compounds at a time in StarDrop, and its speed and performance are just incredible. What used to involve manually wading through thousands of pages of patent applications now takes less than 10 minutes and just a few clicks – simply download the file, open it in StarDrop, run the R-group analysis, and we’re done.
Uwe Heinelt, Medicinal Chemist, Sanofi
Ready to trial StarDrop?
Discover how StarDrop can boost the speed, efficiency, and productivity of your chemistry discovery programmes.
Discover StarDrop Modules
ADME QSAR
High quality predictive QSAR models of a broad range of key ADME and physicochemical properties
Metabolism
Quickly identify metabolic pathways, potential metabolites and guide compound design to avoid metabolic liabilities
Derek Nexus™
Knowledge-based prediction of over 40 key toxicity endpoints, helping guide the design of safe, efficacious drugs
Auto-Modeller™
Build and validate robust QSAR models tailored to your chemistry and data, in an easy and intuitive way
Nova™
Generate and prioritise new, relevant compound ideas using virtual library enumeration and generative chemistry
BIOSTER™
Access a world of chemistry experience, extending Nova’s capabilities with over 29,000 structure modifications and replacement techniques
Inspyra™
Combine your expert chemistry knowledge and the exploratory power of generative methods to identify optimal compounds faster
eSim™ 3D
3D ligand-based design – understand binding conformations to identify and optimise novel active compounds
SeeSAR™
3D structure-based design – visualise ligand and protein structures and identify key interactions driving binding affinity
MPO Explorer™
Develop multi-parameter optimisation strategies using patented rule induction and sensitivity analysis methods
In silico modelling
ADME QSAR
High quality predictive QSAR models of a broad range of key ADME and physicochemical properties
Metabolism
Quickly identify metabolic pathways, potential metabolites and guide compound design to avoid metabolic liabilities
Derek Nexus™
Knowledge-based prediction of over 40 key toxicity endpoints, helping guide the design of safe, efficacious drugs
Auto-Modeller™
Build and validate robust QSAR models tailored to your chemistry and data, in an easy and intuitive way
Generative chemistry
Nova™
Generate and prioritise new, relevant compound ideas using virtual library enumeration and generative chemistry
BIOSTER™
Access a world of chemistry experience, extending Nova’s capabilities with over 29,000 structure modifications and replacement techniques
Inspyra™
Combine your expert chemistry knowledge and the exploratory power of generative methods to identify optimal compounds faster
Linking 2D and 3D SAR and design
eSim™ 3D
3D ligand-based design – understand binding conformations to identify and optimise novel active compounds
SeeSAR™
3D structure-based design – visualise ligand and protein structures and identify key interactions driving binding affinity
MPO Strategy
MPO Explorer™
Develop multi-parameter optimisation strategies using patented rule induction and sensitivity analysis methods
Supporting customisation and collaboration
Real-time collaboration
Accelerate your drug discovery projects by staying aligned on the latest project data, eliminating inefficiencies, and progressing faster together.
Customisation and Integration
Comprehensive data analysis, visualisation and design, easily integrated into your informatics infrastructure.
3D docking and Alignment
Integrated access to 3D modelling to analyse SAR, identify potential liabilities and guide compound design.
Database Access
Seamlessly connect to your in-house databases, with a user-friendly interface to access the latest and most relevant data.
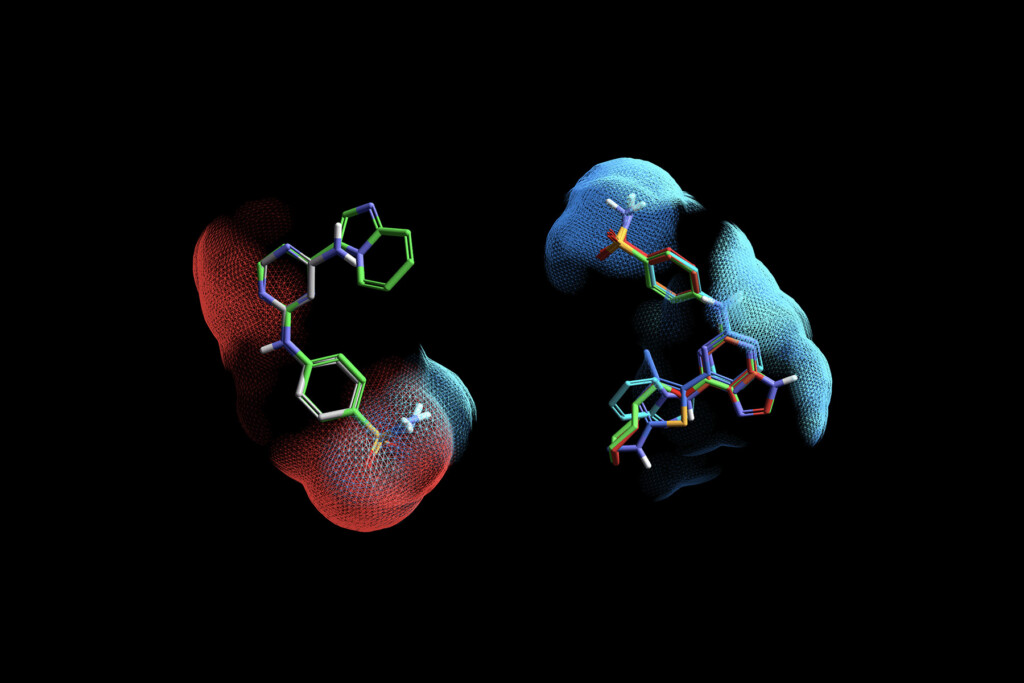
Flexible Deployment
Enjoy identical functionality and user experience, with options for desktop or cloud-based StarDrop, and several hosted server options.
How much will StarDrop cost me?
StarDrop annual subscriptions typically start at around $10,000, but can increase depending on breadth of function and users (e.g. global site licenses with several modules).
Hear from StarDrop users
StarDrop is a very compelling package – multiple powerful tools combined into a single piece of software accessed via a very intuitive user interface. The ease with which powerful MPO scoring functions and QSAR models can be generated and applied to new ideas, as well as the ability to predict metabolic liabilities, make StarDrop the go-to software for ensuring project progression.
Medicinal Chemist, Evotec
We have recently switched to StarDrop from Optibrium and it has been super user friendly.
Co-Founder and Chief Science Officer, Sunflower Wellness
…by integrating StarDrop, IDEAYA can make informed decisions on compound progression, reducing the time and costs associated with drug development.
Head of Research Informatics, IDEAYA
We routinely use StarDrop to visualise complex multi-parameter datasets, which helps inform the next stage of medicinal chemistry design. The user-friendly interface makes data analysis and property prediction easily accessible to medicinal chemists. A great drug discovery tool!
Director, Cerevance
Optibrium tools have proven instrumental during our lead optimization programs and beyond. Particularly, the metabolism module provided valuable guidance on design level during optimization cycles and substantially accelerated structure elucidation during Met-ID studies.
Head of Chemistry, Basilea
We use StarDrop for numerous functions, from the mundane to the elaborate. It’s an incredibly user-friendly, yet powerful piece of software… Our usage spans the entire medicinal chemistry spectrum; from hit triage to the processing of pre-clinical data.
Anima Biotech
StarDrop is a fantastic tool to have as a medicinal chemist, whether you do early-stage medicinal chemistry or deal with a huge amount of data. It’s super quick – I can have a view on my SAR within five minutes, to inform future design strategies.
Medicinal Chemist, adMare Bioinnovations
I really would like to give a big thumbs up to StarDrop and your team. StarDrop is a powerful package tool with an intuitive user interface to access and operate. We really appreciate the visualization module and probabilistic scoring system, which can well-organized interactive charts, graphics and chemical space to facilitate our understanding of the SAR in our chemical series.
Investigator, IFF
The relative ease of use of StarDrop, compared to other drug design and discovery software, allows my students to quickly focus on essential topics, such as ADME, QSAR, metabolism studies, and protein-ligand docking.
Instructor of Chemistry, North Carolina School of Science and Mathematics
Case studies
Super quick SAR analysis
Save time and money
Better collaboration, earlier insights
Ready to trial StarDrop?
Simply complete the form now for a free personalised demo and see how StarDrop fits your unique research needs.
If you’d like to find out more about the StarDrop evaluation process read our step-by-step guide.
