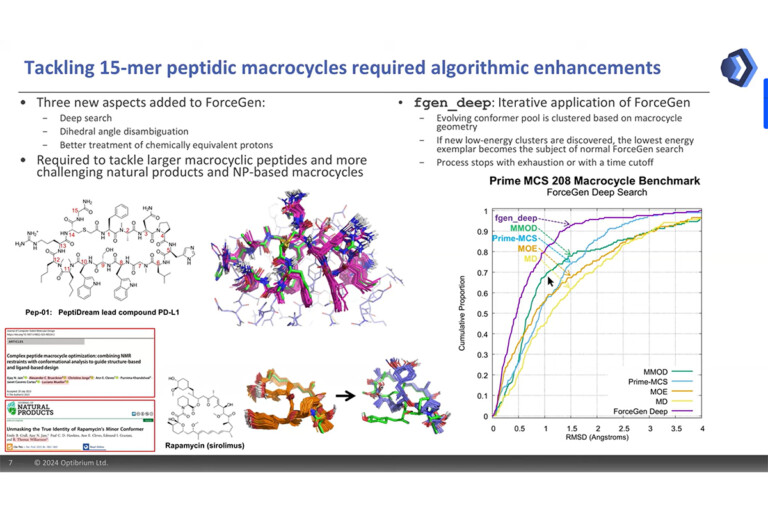
Macrocyclic lead optimisation
Macrocycles are becoming increasingly popular in drug discovery, due to their vast potential against previously “undruggable” targets. But the size…
Visualise your ligands in their protein environment to identify the key interactions driving binding affinity in 3D. These structures can be derived from X-ray crystallography or predicted with any docking software. Additionally, visualise surfaces and H-bond networks to assess the quality of each pose and identify opportunities for optimisation.
Using the award-winning HYDE scoring method, analyse your ligand’s binding affinity with visual atomic contributions and torsion angle heat maps to relate the computational assessments to free energies.
These methods can be applied to docking poses generated using any third-party software, seamlessly linked via the StarDrop Pose Generation Interface.
Generate compound poses for virtual screening and interactive 3D design using FlexX docking. Fast template docking enables you to edit compounds with real-time feedback, while maintaining an established binding mode. Combined with the SeeSAR Affinity module, you can assess the impact of new optimisations on binding energetics.
Macrocycles are becoming increasingly popular in drug discovery, due to their vast potential against previously “undruggable” targets. But the size…

From the manuscript “DOCKSTRING: Easy Molecular Docking Yields Better Benchmarks for Ligand Design”, Miguel García-Ortegón, Sergio Bacallado, et al¹ have developed…

This worked example explores ways to assess and design compounds in 3D using the SeeSAR Pose module.
The SeeSAR modules are available as optional modules for StarDrop. To try StarDrop please complete the form and a member of the team will get in touch to understand your needs.