Enhanced ADME predictions : New and improved models in StarDrop 8
All models are available in StarDrop’s ADME QSAR module: pKa– Most acidic pKa– Most basic pKa– All pKa valueslogP (Octanol/Water)logD7.4 (Octanol/buffer…

All models are available in StarDrop’s ADME QSAR module: pKa– Most acidic pKa– Most basic pKa– All pKa valueslogP (Octanol/Water)logD7.4 (Octanol/buffer…

The importance of predicting toxicity early in drug development Highlighting safety and toxicity early in the drug development process is…

How can I predict my compound’s absorption? The first of the ADMET properties relate to absorption. Understanding how a drug…

What are StarDrop and Semeta? Semeta is a tailored platform for DMPK scientists. It enables users to address key challenges…

Data curation for model building A model can only be as good as the data it has been trained on.…

Why focus on cytochrome P450 enzymes? CYPs are a ubiquitous superfamily of heme-containing monooxygenases responsible for approximately 70–80% of observed…

What’s the purpose of a predictive model? What’s the value of predictive models for drug discovery? Most of the undergraduate…

Discover which metabolite prediction software is best for your needs in this comprehensive guide from Optibrium. Compare top tools like Meteor Nexus, MetaSite, and StarDrop to make informed decisions for drug metabolism prediction

How number of users affect drug discovery software costs The number of people who need access to the platform is…

Peer-reviewed study published in Xenobiotica describes an innovative new method that predicts the routes and products of Phase I and II metabolism with high sensitivity and greater precision than
other approaches

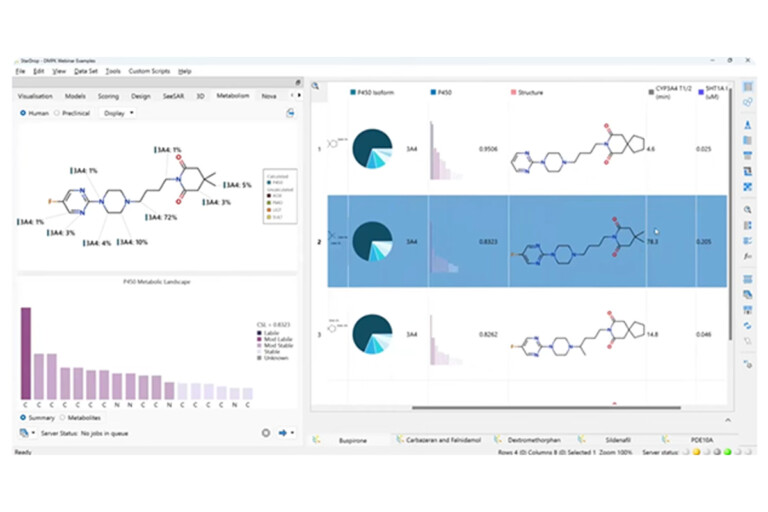
In this example we will explore the feasibility of pursuing a fast-follower for Buspirone, a 5-HT1A ligand used as an anti-anxiolytic therapeutic, which has a known liability due to rapid metabolism by CYP3A4.
Interpreting metabolite-ID experiments; determining the right species for animal studies; providing optimisation suggestions for your medicinal chemistry colleagues to overcome…
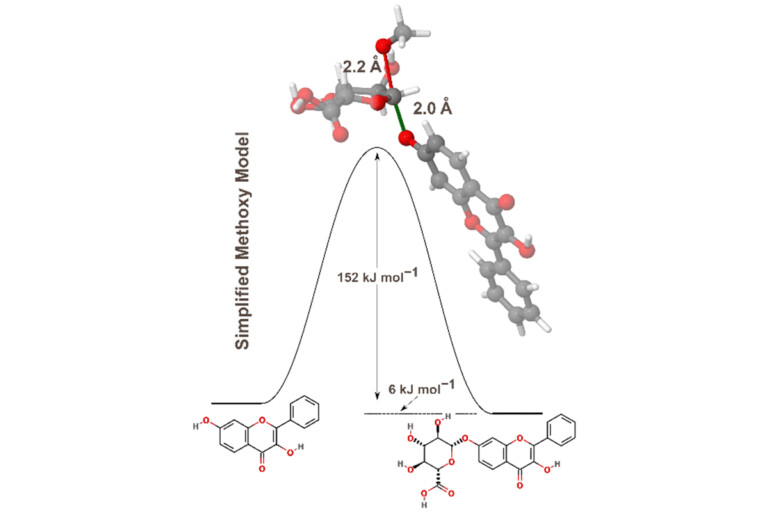
Predicting metabolism at an early stage is important in maximising the chance of a drug’s success. However, accurate, useful models…
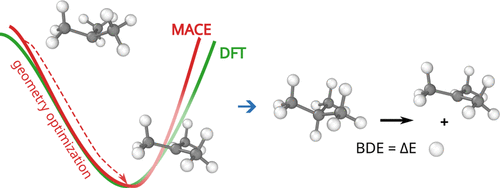
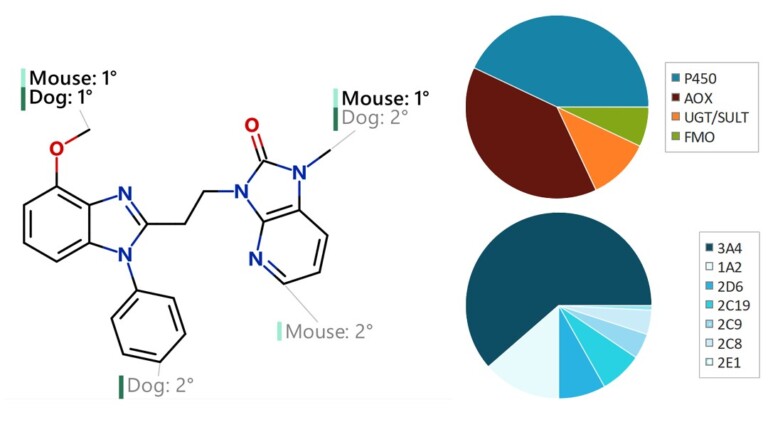
This peer-reviewed paper in Xenobiotica describes a new method to determine the most likely experimentally-observed routes of metabolism and metabolites based on our WhichP450™, regioselectivity and new WhichEnzyme™ model.
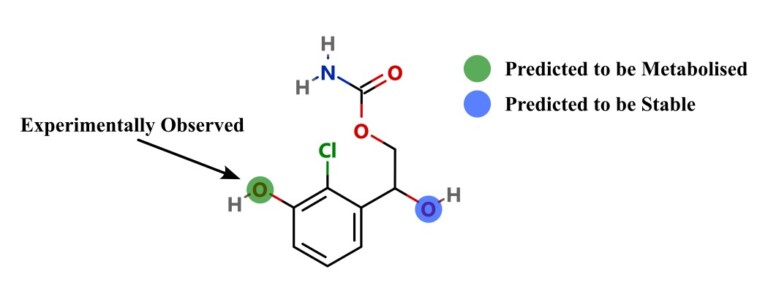
This paper describes a model to predict whether a particular site on a molecule will be metabolised by cytosolic sulfotransferase enzymes (SULTs).
Watch Optibrium CEO Matt Segall and Principal Scientist Mario Öeren as they explore groundbreaking new quantum mechanics and machine learning models which go beyond P450s and provide insights on a broad range of enzymes involved in drug metabolism.
Introduction Predicting sites of metabolism (SoM) enable chemists to be more efficient in optimising the structure of new chemical entities…
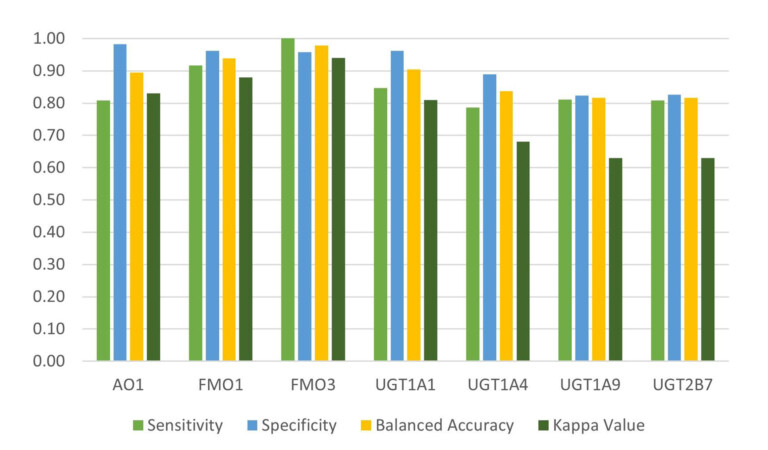
This paper describes the prediction of the regioselectivity of metabolism by AOs, FMOs and UGTs for humans and CYPs for three preclinical species.
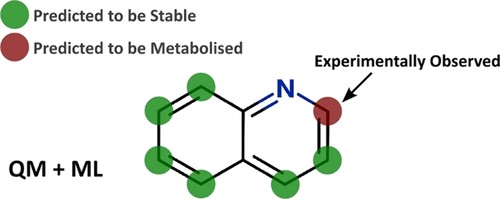
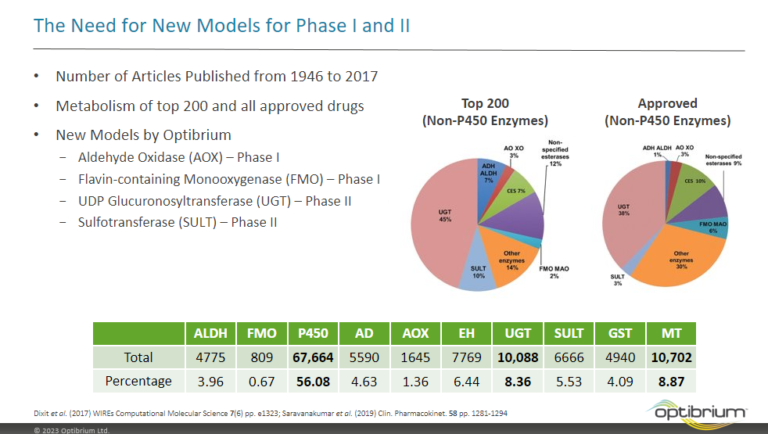
Methods for modelling two enzyme families, flavin-containing monoxygenases (FMOs) and uridine 5′-diphospho-glucuronosyltransferases (UGTs), to predict reactivity to drug metabolism.