This new peer-reviewed paper in the Journal of Chemical Information and Modeling describes a model to predict whether a particular site on a molecule will be metabolised by cytosolic sulfotransferase enzymes (SULTs). The paper builds on previous metabolism prediction research, developing the capabilities available in StarDrop and Semeta.
Summary
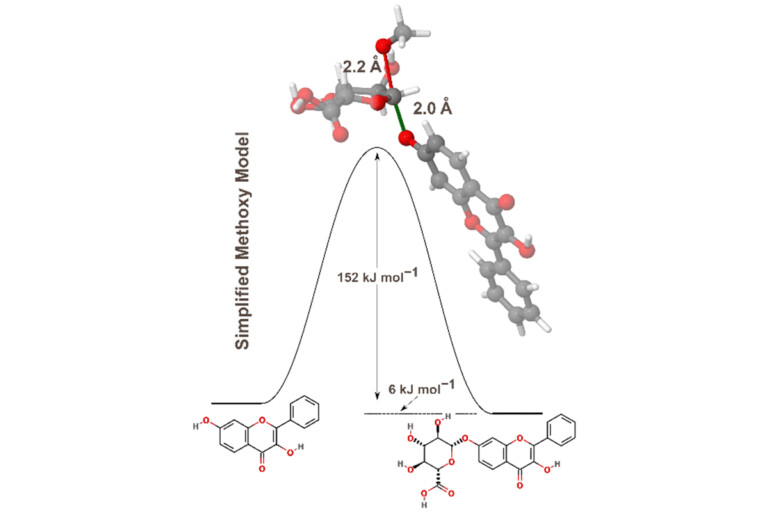
During Phase II drug metabolism, also known as the conjugation phase, compounds are conjugated to other polar compounds. The main enzymes involved are uridine 5′-diphospho-glucuronosyltransferases (UGTs). However another prominent family during this phase are cytosolic sulfotransferases (SULTs). These enzymes are responsible for sulfation reactions. To discover how small molecules will be metabolised in this conjugation phase, it is therefore crucial to see how the regioselectivity of SULTs differ from that of UGTs.
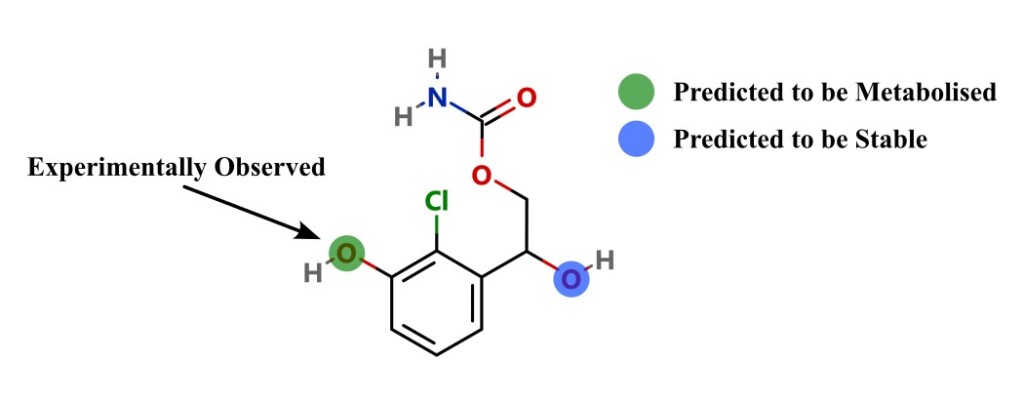
In this paper, the research team developed a general ligand-based model, to predict whether a site will be metabolized by SULTs. During the study, they found that the SULT binding site was the main influence on regioselectivity of metabolism, rather than any influence by the activation energy of the rate-limiting step of the catalysis. Therefore, they trained the model only on steric and orientation descriptors. This mimicked the SULT binding pocket, generating a model with very good accuracy values.
Find out more about predicting drug metabolism
If you would like to discover more of our metabolism research, then take a look at our J. Med. Chem. paper, which describes how to predict the regioselectivity of AO, CYP, FMO and UGT metabolism. Alternatively, watch our webinar with authors Mario Öeren and Matthew D. Segall on the integrated prediction of Phase I and II metabolism.
More drug metabolism resources

Private: Integrated AI synthesis prediction in drug discovery
Learn more about how AI, machine learning and other computational tools can support the discovery process, bringing you feasible synthetic routes to your target compounds.

Predicting reactivity to drug metabolism: beyond P450s – modelling FMOs and UGTs
Methods for modelling two enzyme families, flavin-containing monoxygenases (FMOs) and uridine 5′-diphospho-glucuronosyltransferases (UGTs), to predict reactivity to drug metabolism.