To start a virtual screening experiment, click the arrow button menu  Run Virtual Screen.
Run Virtual Screen.

Several options are available to load the 3D input structure(s) to use for virtual screening:
Open a file containing the 3D coordinates by clicking the  button.
button.
Download a PDB containing the molecule in a bioactive conformation by clicking the  button.
button.
Use an entire binding hypothesis or obtain a molecule in a specific conformation from a binding hypothesis result by clicking the  button.
button.
Use a compound in the 3D Viewer by selecting all atoms of the molecule and clicking the  button.
button.
Once the desired input conformations have been selected, click Next.
In the next page of the wizard, you can Choose a virtual screening database.
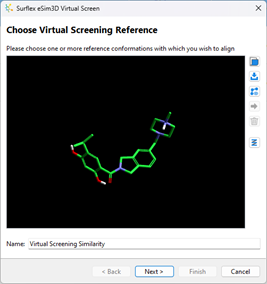
Load the virtual screen collection by clicking on  button, locate where you saved the files and confirm it.
button, locate where you saved the files and confirm it.
For more details, please refer to the StarDrop user guide.