This paper describes the prediction of the regioselectivity of metabolism by AOs, FMOs and UGTs for humans and CYPs for three preclinical species. The research extends from previous work developing the StarDrop Metabolism module. Visit the Metabolism webpage to discover how StarDrop can help you model metabolism.
Summary
Predicting the sites of metabolism for different key enzymes across the various phases of drug metabolism is vital in avoiding the failure of many late-stage drug candidates or even withdrawal of approved drugs. This study presents methods for predicting the isoform-specific metabolism for several key enzymes involved in drug metabolism across both Phase I (modification) and II (conjugation). These include human aldehyde oxidases (AOs), flavin-containing monooxygenases (FMOs) and uridine 5′-diphospho-glucuronosyltransferases (UGTs). General models of cytochrome P450 (CYP) metabolism for preclinical species were also developed.
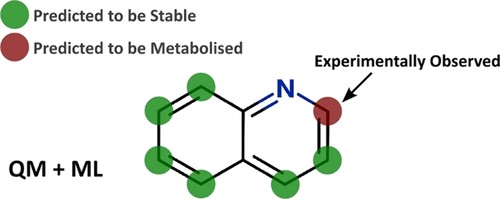
Using semi-empirical quantum mechanical simulations, the reactivity of each site of metabolism is evaluated holistically in the context of the whole molecule. This reactivity information is then combined with the orientation and steric effects of the binding pockets of the different enzyme isoforms, to generate predictive models, achieving excellent balanced accuracy and kappa values.
Citation Details
M. Öeren, P. J. Walton, J. Suri, D. J. Ponting, P. A. Hunt, and M. D. Segall, J. Med. Chem. 2022, 65, 20, 1406–1408. DOI: 10.1021/acs.jmedchem.2c01303
Find out more
If you would like to discover more of our metabolism research, take a look at our recent J. Chem. Inf. Model. paper, which describes how to predict the regioselectivity of SULT metabolism, or watch our webinar on the integrated prediction of Phase I and II metabolism.