From X-ray refinement to lead optimisation
Conventional ligand-fitting and refinement methods in X-ray electron density maps often yield models with unrealistically high conformational strain. This occurs because ligands are typically treated as single conformers using atom-specific B-factors. These parameters often introduce geometric distortions or obscure biologically meaningful pose variations that might misguide lead optimisation.
xGen™ is a real-space refinement method that addresses these problems by balancing how well a ligand fits the electron density with its internal conformational strain. Rather than a single static pose, it produces a realistic ensemble of conformers that collectively explain the experimental data. This approach reveals meaningful structural variations and yields physically plausible models with substantially reduced strain.
The xGen method
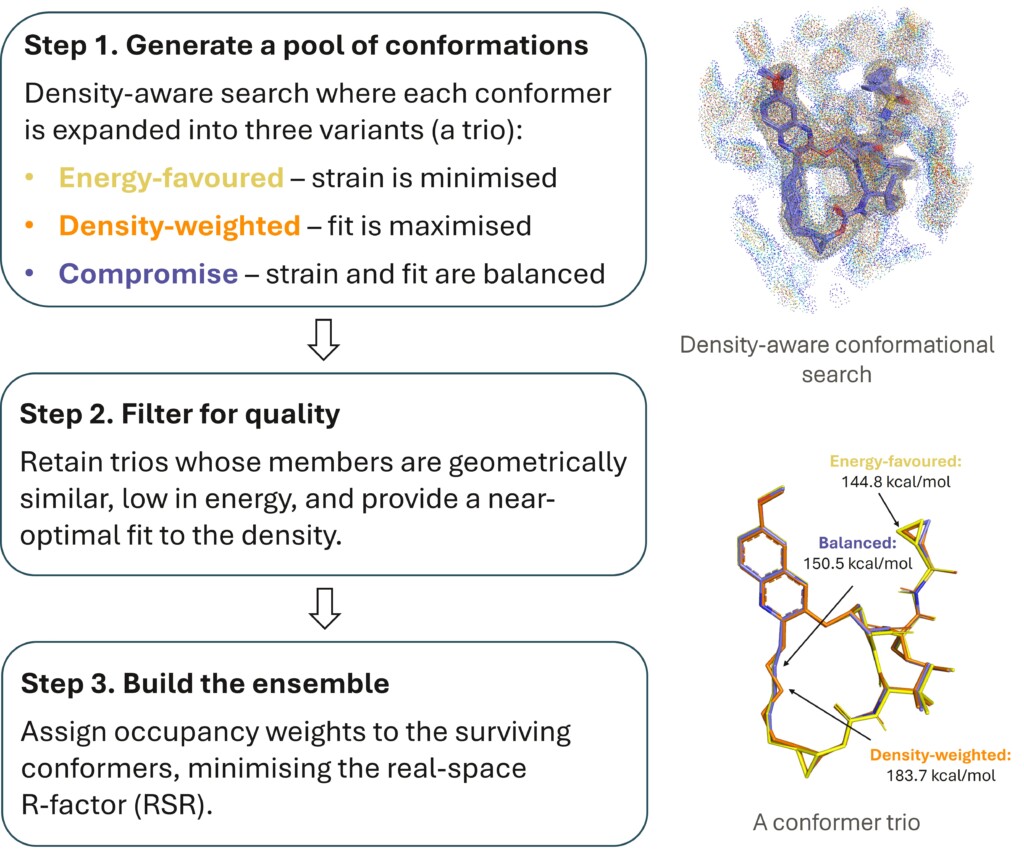
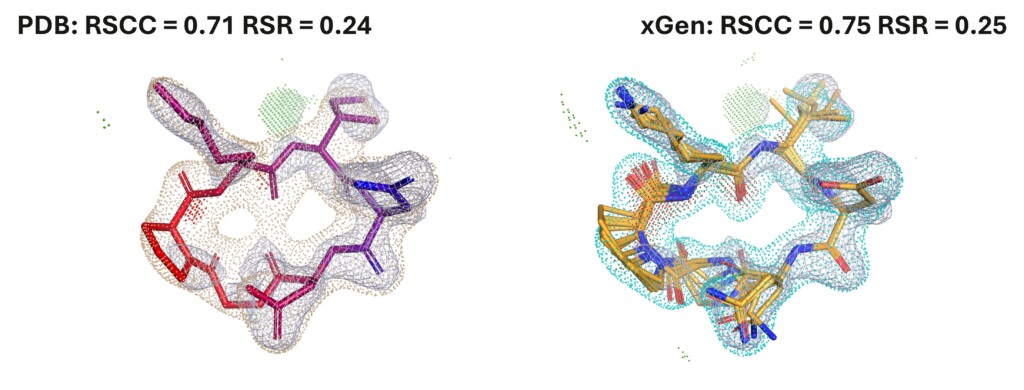
xGen ensembles reduce ligand strain by half
Validated on ~3,000 complexes, xGen ensembles reduce artificially high PDB ligand strain by ~50% while improving density fits. This occupancy-weighted approach provides physically realistic, low-energy alternatives to traditional single-conformer models.
| Average strain (kcal/mol) | Deposited | xGen |
| Macrocycles | 6.8 | 3.7 |
| Non-macrocycles | 4.2 | 2.5 |
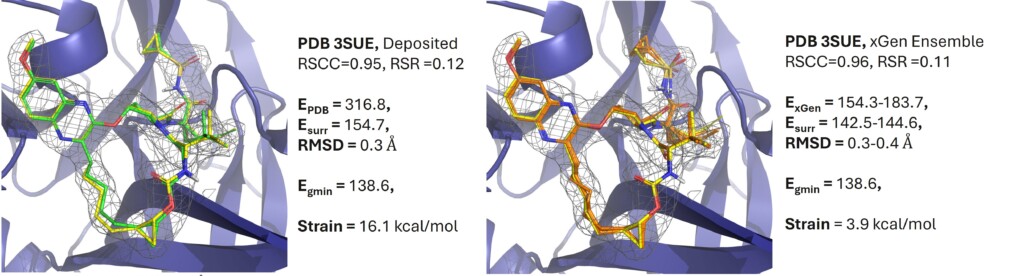
Reduce synthetic effort by up to 90%
In an integrated workflow, xGen-generated low-strain ensembles of the lead peptide were used to derive conformational restraints. Combined with docking and scoring, this approach:
- accurately ranked compounds by predicted binding energies,
- generated correct poses for docked macrocycles.
The conformational strain emerged as the dominant predictor of ligand binding, and the clinical candidate was identified in the top-ranked 10% of synthesised analogues.
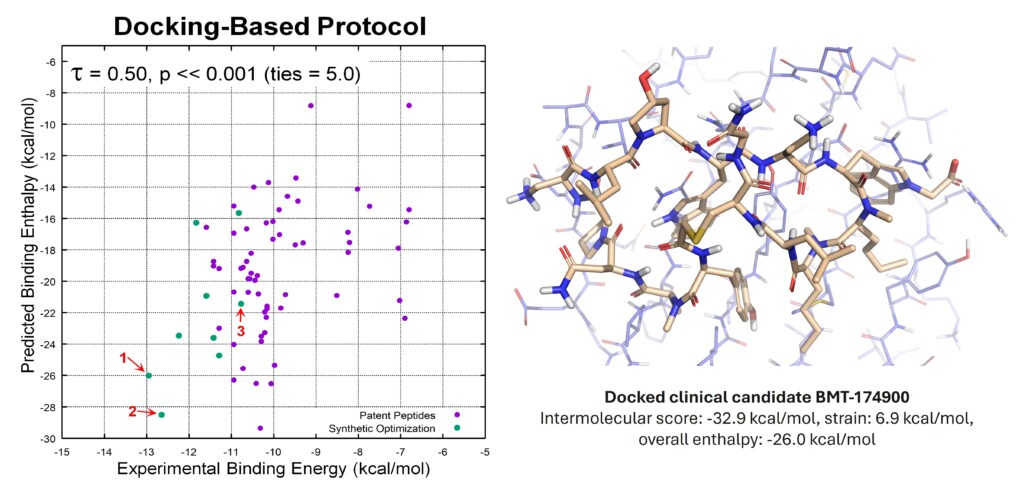
The importance of strain
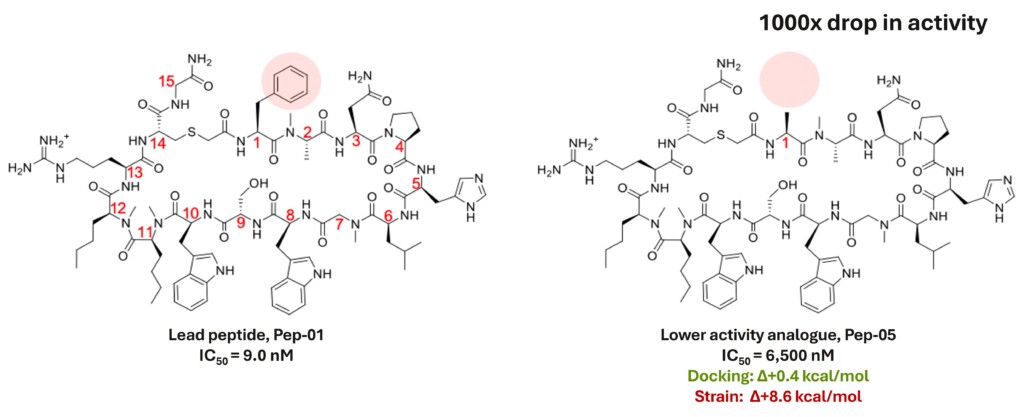
Conclusions
- xGen offers a paradigm shift for ligand modelling, producing physically realistic conformer ensembles for ligands with lower strain.
- Ligand strain is a predictive factor for macrocyclic peptide optimisation: If the ligand has high strain, aim to optimise its geometry, and if it already has low strain, improve protein-ligand interaction footprint.
- Accurate pose prediction enables effective CADD-driven macrocycle design, boosting efficiency and productivity.
References
- Jain, A. N. et al. xGen: Real-Space Fitting of Complex Ligand Conformational Ensembles to X‑ray Electron Density Maps. J. Med. Chem. 2020, 63, 10509-10528
- Jain, A. N. et al. Complex peptide macrocycle optimization: combining NMR restraints with conformational analysis to guide structure‑based and ligand‑based design. JCAMD. 2023, 37, 519-535
Acknowledgements
- Merck: Alexander C. Brueckner, Mikhail Reibarkh, and Edward C. Sherer
- Optibrium: James Halle and Nathan Brown