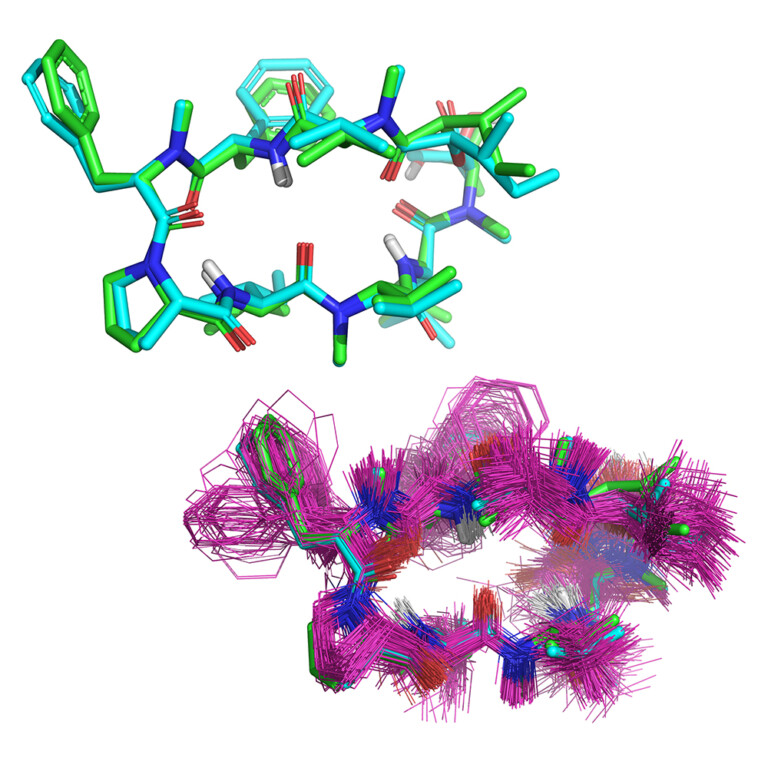
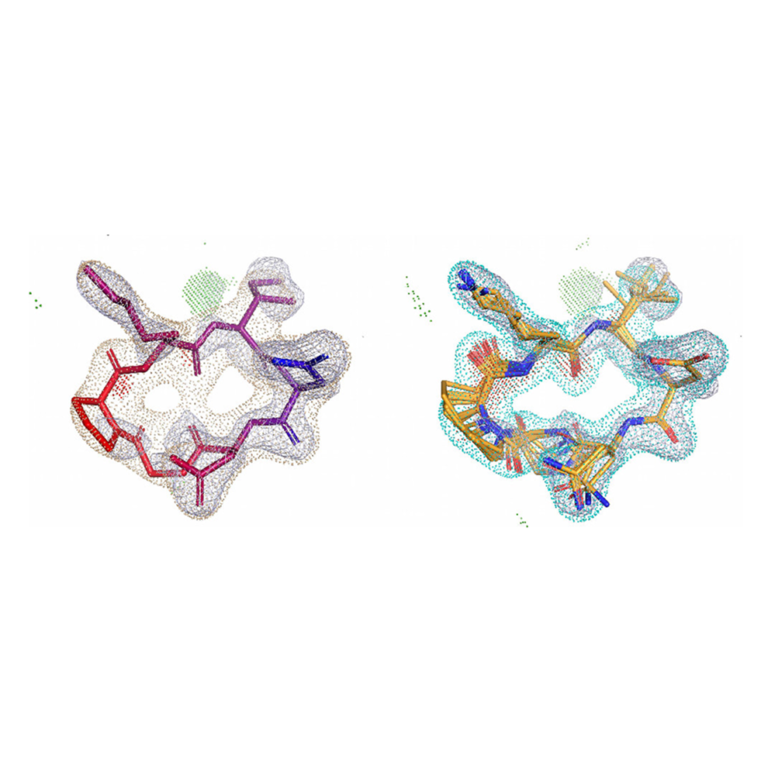
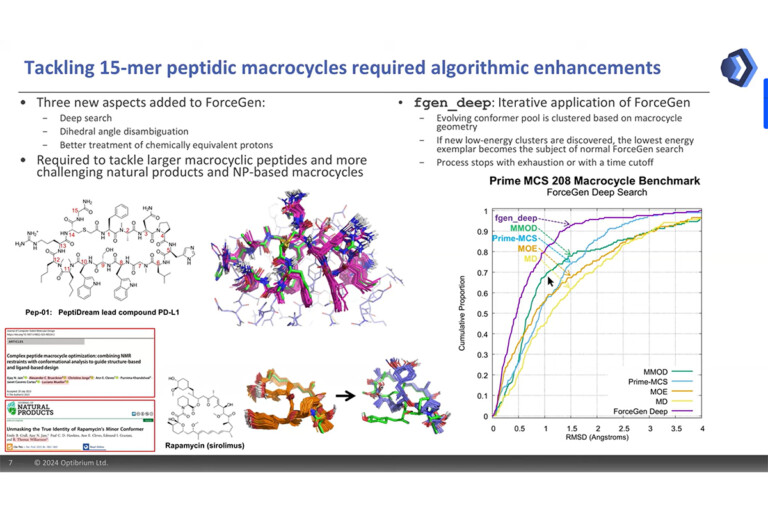
Access top-tier docking enrichment and accurate pose prediction with reliable automated docking procedures. Automate protein preparation and reduce inappropriate bias with Surflex-Dock’s easy-to-use workflows. Surflex-Dock supports your 3D structure-based design efforts, allowing you to model ligands from small molecules to large macrocycles.
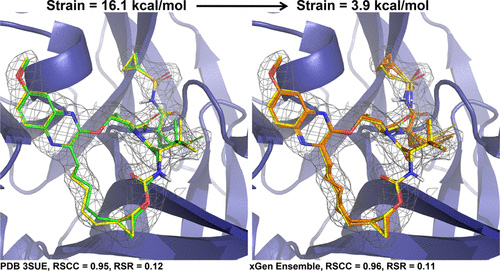
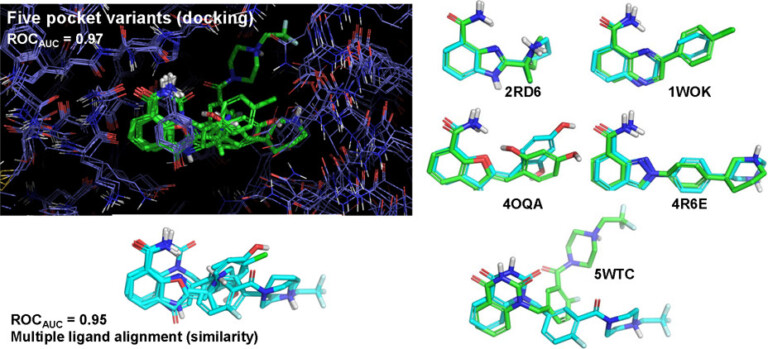
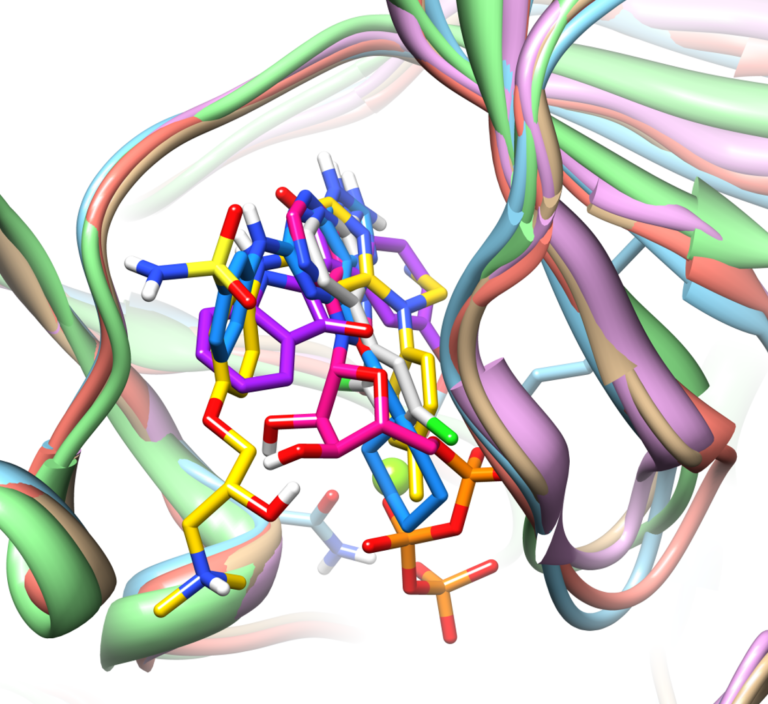
With Surflex-Dock, ensembles of protein conformations take into account the variability in the binding pocket and relevant interactions, to give a more realistic picture of molecular docking.
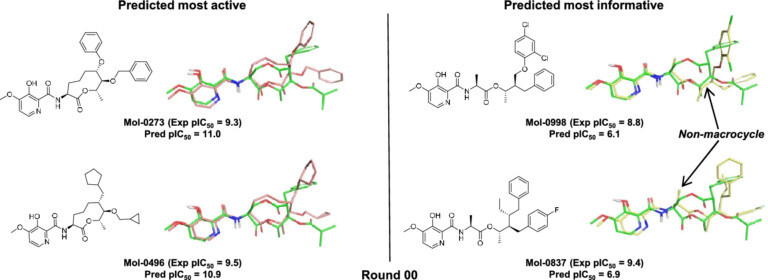
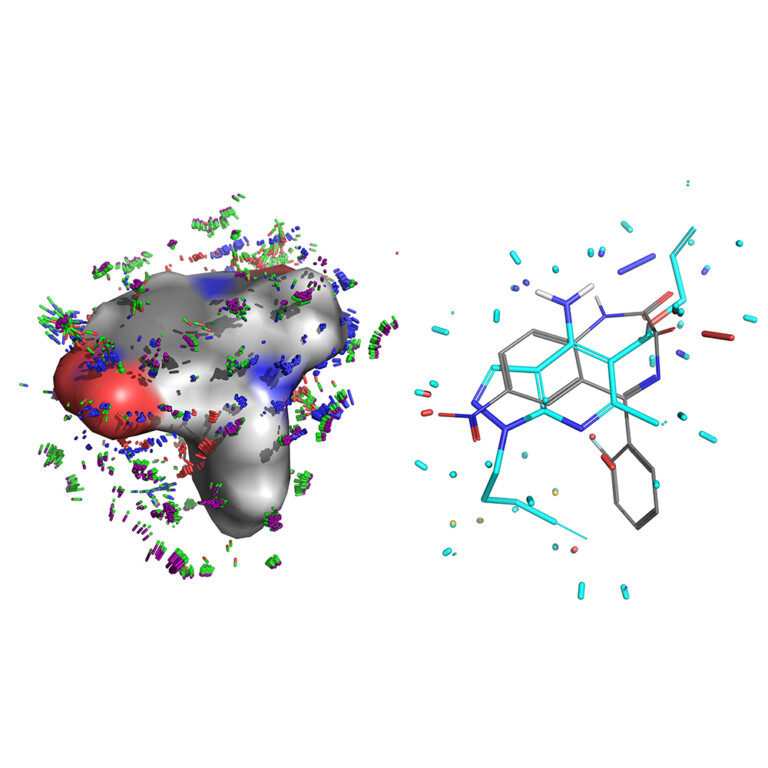
Surflex-Dock enables highly accurate non-cognate ligand docking, so you can correctly predict poses for future potential ligands for your protein complex, making use of known prior bound ligand poses with eSim™.
Surflex-Dock predictions can be accessed through our intuitive and highly visual PyMOL GUI.